FDA
With CORE, FDA brought together expertise in medicine, public health and science to coordinate its efforts to find, stop, and prevent foodborne illness outbreaks. Since CORE was established in 2011, CORE teams have identified 959 potential outbreaks, responded to 234 outbreaks potentially linked to FDA regulated food products, identified a specific food in 100 outbreaks, and warned consumers to avoid those foods through more than 400 public notifications.
Outbreak Detection, Response, Prevention
On the Lookout
The CORE Signals and Surveillance Team evaluates emerging outbreaks and disease surveillance trends, working in collaboration with CDC, FDA field offices, and state agencies. The team reviews firm data including past inspections, sampling results, product distribution, and sourcing information. It also considers previous incidents involving similar pathogen and food pairs. This information is used to determine whether it can provide clues to understand emerging outbreaks. When an outbreak appears to be caused by an FDA-regulated food, this information is passed to a Response Team to coordinate FDA’s response efforts.
On the Hunt
Response Teams have one goal: to control and stop the outbreak. Response Teams work directly with FDA field offices, FDA subject-matter experts, the CDC, and state partners on a response strategy. The team coordinates investigations, inspections, sampling, and traces product distribution. Close coordination among the FDA, CDC, and state and local regulatory, public health and agriculture departments is crucial to stopping an outbreak.
Results of Response Activities
During or following an outbreak response several actions can be taken to either protect public health or inform public health efforts. Among the actions that have been taken as a result of CORE-coordinated investigations are:
- More than 400 Public Advisories since 2011
- At least 251 Recalls, including downstream recalls since 2011
- 268 CORE-issued Assignments, including food facility/farm investigations/inspections, record collection, and sample collection related to outbreaks between 2016 – 2019
- 106 Assignments with sample collection between 2016 – 2019
Communications
The CORE Communications Team monitors emerging and active incident investigations. If there is an ongoing risk to the public and actionable steps can be taken to reduce risk of illness, the FDA will issue public warning. This team also prepares responses to inquiries from FDA stakeholders and the media regarding outbreaks.
An Eye to Prevention
What did we learn? How can we prevent this from happening again? These questions guide the mission of the Outbreak Evaluation and the Outbreak Analytics Teams. These teams look at all aspects of the outbreak, from ingredient sourcing to production and distribution. They conduct data analyses to recommend ways to integrate preventative measures in food safety activities.
Results of Post-Response Activities
The CORE Outbreak Evaluation and Outbreak Analysis Teams have used data from CORE-coordinated outbreaks to contribute to:
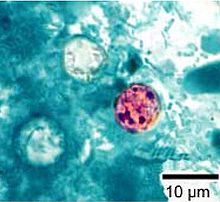
- The development of improved detection of the Cyclospora parasite in foods to improve outbreak detection and prevention efforts.
- The development of the FDA Produce Safety Rule, aimed at reducing the risk of contamination of produce, and related documents.
- The development of inspectional and sampling surveillance assignments to monitor firms and industries with foods associated with outbreaks and gather outbreak prevention data.
- Providing resources to retailers, growers, shippers, and carriers on handling produce recalled after an outbreak and develop articles and presentations focused on past outbreak investigations to inform and educate the public and food industry professionals.
- Communicating the results of outbreak analyses and prevention efforts through scientific journal articles and professional conferences focused on outbreak response and prevention.
More Information