Following the investigations carried out by the Belgian health authorities, together with their English, European and in particular French counterparts, the company Ferrero proceeded on April 5, 2022 to the recall of several Kinder range products manufactured in a factory in Belgium due to suspected contamination by Salmonella Typhimurium . On April 8, 2022, the recall finally affected all Kinder products from this factory, regardless of their expiry date. On April 14, 2022, an update of the recalled products, including the 2021 Christmas Advent Calendars, was released.
Case of salmonellosis in France: update on June 2, 2022
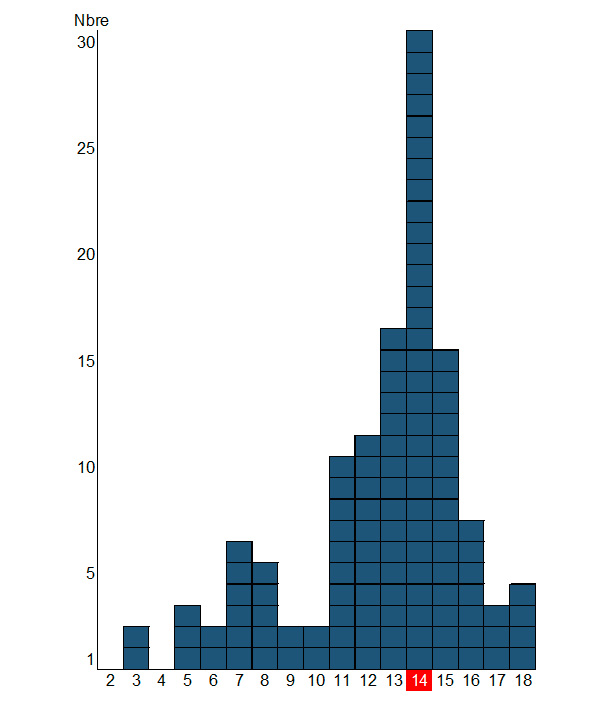
In total, as of 02/06/2022: 118 cases of salmonellosis with a strain belonging to the epidemic have been identified by the National Reference Center (CNR) for salmonella at the Institut Pasteur in France (figure 1) .
Figure 1 – Epidemic curve: number of confirmed cases of salmonellosis caused by Salmonella Typhimurium, monophasic variant (cluster 1 HC5_296366 and cluster 2 HC5_298160), by week of isolation (with in red the week corresponding to the recall of products from the production plant) ‘Arlon in Belgium) – Metropolitan France, weeks 2 to 18, 2022 (N=118)
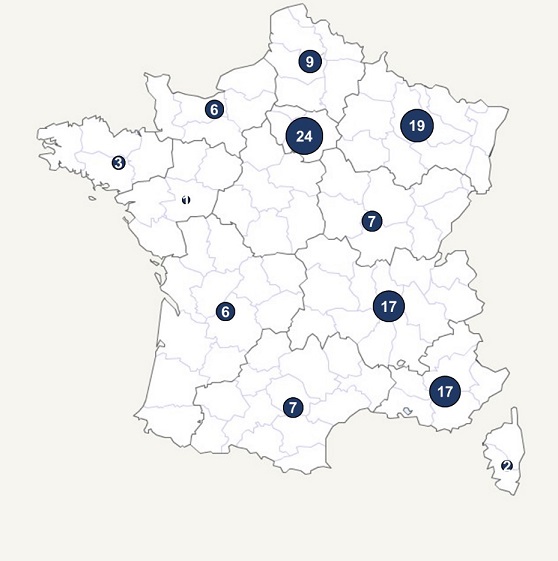
The 118 cases are spread over 12 metropolitan regions (Ile-de-France (24 cases), Grand-Est (19 cases), Auvergne-Rhône-Alpes (17 cases), Provence-Alpes-Côte d’Azur (17 cases) , Hauts-de-France (9 cases), Bourgogne-Franche-Comté (7 cases), Occitanie (7 cases), Normandy (6 cases), New Aquitaine (6 cases), Brittany (3 cases), Corsica (2 cases) and Pays de la Loire (1 case)) with a median age of 4 years, and concern 57 girls and 61 boys.
Figure 2 – Geographical distribution of confirmed cases of salmonellosis due to Salmonella Typhimurium, monophasic variant (cluster 1 HC5_296366 and cluster 2 HC5_298160), by region of residence – metropolitan France, weeks 2 to 18, 2022
Fifty-one cases were questioned by Public Health France. All the cases, except 1, report, before the onset of their symptoms (which occurred between 20/01 and 04/04/2022), the consumption of chocolates of the brand cited here.
Twenty-two people were hospitalized for their salmonellosis, all since discharged. No deaths were reported.
The foods in question having been identified and the management measures taken, the weekly situation updates are drawn up. Public Health France continues to monitor the reporting of cases by the NR, which are expected due to the different delays inherent in monitoring ( see the infographic dedicated to food alerts ).
The successive withdrawals and recalls of the Kinder brand products concerned, produced by the Belgian factory with its closure by the Belgian authorities, should limit the occurrence in France of new cases of salmonellosis in connection with these chocolates.
The possible identification of new cases with dates of isolation at a distance from the recall withdrawal measures will be the subject of investigations if necessary.
To find out the list of products concerned by the withdrawal-recall: https://rappel.conso.gouv.fr/
People who have consumed the products mentioned above and who present symptoms (gastrointestinal disorders, fever within 72 hours of consumption), are invited to consult their doctor without delay, notifying him of this consumption.
In order to limit person-to-person transmission (especially in households with young children), it is recommended to wash your hands well with soap and water after using the toilet, after changing your child, and before to cook.