Archives
-
Join 346 other subscribers
KSWFoodWorld
Blog Stats
- 445,811 Views
Category Archives: Microbiological Risk Assessment
Research – Bacteriological Quality and Biotoxin Profile of Ready-to-Eat Foods Vended in Lagos, Nigeria
A comprehensive study of bacterial and biotoxin contaminants of ready-to-eat (RTE) foods in Nigeria is yet to be reported. Hence, this study applied 16S rRNA gene sequencing and a dilute-and-shoot LC-MS/MS method to profile bacteria and biotoxins, respectively, in 199 RTE food samples comprising eko (n = 30), bread (n = 30), shawarma (n = 35), aadun (n = 35), biscuits (n = 34), and kokoro (n = 35). A total of 631 bacterial isolates, clustered into seven operational taxonomic units, namely Acinetobacter, Bacillus, Klebsiella, Proteus and Kosakonia, Kurthia, and Yokenella, that are reported for the first time were recovered from the foods. One hundred and eleven metabolites comprising mycotoxins and other fungal metabolites, phytoestrogenic phenols, phytotoxins, and bacterial metabolites were detected in the foods. Aflatoxins, fumonisins, and ochratoxins contaminated only the artisanal foods (aadun, eko, and kokoro), while deoxynivalenol and zearalenone were found in industrially-processed foods (biscuit, bread, and shawarma), and citrinin was present in all foods except eko. Mean aflatoxin (39.0 µg/kg) in artisanal foods exceeded the 10 µg/kg regulatory limit adopted in Nigeria by threefold. Routine surveillance, especially at the informal markets; food hygiene and safety education to food processors and handlers; and sourcing of high-quality raw materials are proposed to enhance RTE food quality and safeguard consumer health.
Posted in Aflatoxin, Aspergillus Toxin, Decontamination Microbial, Food Micro Blog, Food Microbiology, Food Microbiology Blog, Food Microbiology Research, Food Microbiology Testing, Food Toxin, Fusarium Toxin, microbial contamination, Microbial growth, Microbiological Risk Assessment, Microbiology, Microbiology Investigations, Microbiology Risk, Mold Toxin, Mould Toxin, Mycotoxin, Toxin
Research – Current Perspectives on Viable but Non-Culturable Foodborne Pathogenic Bacteria: A Review
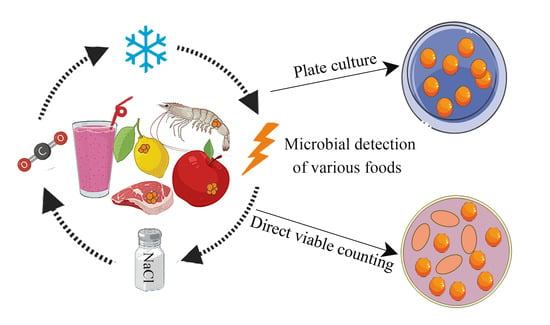
Abstract
Foodborne diseases caused by foodborne pathogens pose risks to food safety. Effective detection and efficient inactivation of pathogenic bacteria has always been a research hotspot in the field of food safety. Complicating these goals, bacteria can be induced to adopt a viable but non-culturable (VBNC) state under adverse external environmental stresses. When in the VBNC state, pathogens cannot form visible colonies during traditional culture but remain metabolically active and toxic. The resulting false negative results in growth-related assays can jeopardize food safety. This review summarizes the latest research on VBNC foodborne pathogens, including induction conditions, detection methods, mechanism of VBNC formation, and possible control strategies. It is hoped that this review can provide ideas and methods for future research on VBNC foodborne pathogenic bacteria.
Posted in Food Micro Blog, Food Microbiology, Food Microbiology Blog, Food Microbiology Research, Food Microbiology Testing, Food Pathogen, microbial contamination, Microbial growth, Microbiological Risk Assessment, Microbiology, Microbiology Investigations, Microbiology Risk, Pathogen, pathogenic, Research
Research – Detection of Escherichia coli O157:H7 in imported meat products from Saudi Arabian ports in 2017
Escherichia coli O157:H7 is a foodborne pathogen, which causes various health conditions in humans, including fatigue, nausea, bloody diarrhoea and in some cases, even death. In 2017, 15.71% of the total imported food products in Saudi Arabia (SA) were meat-based. India and Brazil are two of the top five countries from where SA imports meat. According to the Saudi Food and Drug Authority, in 2017, at least 562, 280, and 50 samples of imported beef, chicken and sheep meat, respectively, were tested for the presence of E. coli O157:H7. Amongst these, E. coli O157:H7 was detected in respectively 6.80% and 2.20% of the tested beef meat samples imported from India and Brazil as well as in respectively 6.96% and 3.57% of the tested chicken samples imported from Brazil and Ukraine. Moreover, the pathogen was detected in 2.13% of the tested sheep meat samples imported from India. The present report provides evidence that imported meat can serve as the carrier of E. coli O157:H7, which may lead to epidemics within the Kingdom of Saudi Arabia.
Posted in E.coli O157, E.coli O157:H7, Food Micro Blog, Food Microbiology, Food Microbiology Blog, Food Microbiology Research, Food Microbiology Testing, microbial contamination, Microbial growth, Microbiological Risk Assessment, Microbiology, Microbiology Investigations, Microbiology Risk, Research, STEC, STEC E.coli
Research – Heterotrophic Plate Count Can Predict the Presence of Legionella spp. in Cooling Towers
Abstract
Legionella pneumophila (Lp) colonizes aquatic environments and is a potential pathogen to humans, causing outbreaks of Legionnaire’s disease. It is mainly associated with contaminated cooling towers (CTs). Several regulations, including Spanish legislation (Sl), have introduced the analysis of heterotrophic plate count (HPC) bacteria and Legionella spp. (Lsp) in management plans to prevent and control Legionella outbreaks from CTs. The 2003 Sl for CTs (RD 865/2003) considered that concentrations of HPC bacteria ≤10,000 cfu/mL and of Lsp ≤100 cfu/L are safe; therefore, no action is required, whereas management actions should be implemented above these standards. We have investigated to what extent the proposed standard for HPC bacteria is useful to predict the presence of Lsp in cooling waters. For this, we analyzed Lsp and HPC concentrations, water temperature, and the levels of chlorine in 1376 water samples from 17 CTs. The results showed that in the 1138 water samples negative for Legionella spp. (LN), the HPC geometric mean was significantly lower (83 cfu/mL, p < 0.05) than in the positive Lsp. samples (135 cfu/mL). Of the 238 (17.3%) LP samples, 88.4% (210/238) were associated with values of HPC ≤10,000 cfu/mL and most of them showed HPC concentrations ≤100 (53.7%). In addition, a relatively low percentage of LP (28/238, 11.6%) samples were associated with HPC bacteria concentrations >10,000 cfu/mL, indicating that this standard does not predict the colonization risk for Legionella in the CTs studied. The present study has demonstrated that a threshold concentration ≤100 cfu/mL of HPC bacteria could better predict the higher concentration of Legionella in CTs, which will aid in preventing possible outbreaks.
Posted in Cooling Towers, Decontamination Microbial, Food Micro Blog, Food Microbiology, Food Microbiology Blog, Food Microbiology Research, Food Microbiology Testing, Legionella, Legionnaires’ disease, microbial contamination, Microbial growth, Microbiological Risk Assessment, Microbiology, Microbiology Investigations, Microbiology Risk, Water, water microbiology, Water Safety
Research – Effect of High Hydrostatic Pressure Processing on the Microbiological Quality and Bacterial Diversity of Sous-Vide-Cooked Cod
Abstract
High hydrostatic pressure (HP) is a promising method to improve the microbiological quality of sous-vide foods. Monitoring the composition and behavior of the microbial communities in foods is of most importance for the production of high-quality and safe products. High-throughput sequencing (HTS) provides advanced approaches to determine food’s microbial community composition and structure. The aim of the present study was to determine the impact of different HP treatments on the microbial load and bacterial diversity of sous-vide Atlantic cod. Sous-vide cooking at 57.1 °C for 30 min followed by HP treatment at 500 MPa for 8 min reduced viable cell counts (total aerobic mesophiles) in the cod samples below detectable levels for 45 days of storage under refrigeration. In a second trial with cod cooked sous-vide at 52 °C for 20 min followed by HP treatments at 300 or 600 MPa (with HP treatment temperatures of 22 °C or 50 °C for 4 or 8 min, depending on treatment), only the treatments at 600 MPa delayed bacterial growth for at least 30 days under refrigeration. The optimal HP conditions to improve the microbiological quality of sous-vide cod cooked at low temperatures were obtained at 600 MPa for 4 min at a pressurization temperature of 50 °C. Bacterial diversity was studied in cod cooked sous-vide at 52 °C for 20 min by HTS. In the absence of HP treatment, Proteobacteria was the main bacterial group. A succession of Pseudomonadaceae (Pseudomonas) and Enterobacteriaceae was observed during storage. Firmicutes had low relative abundances and were represented mainly by Anoxybacillus (early storage) and Carnobacterium (late storage). The HP-treated sous-vide cod showed the greatest differences from controls during late storage, with Aerococcus and Enterococcus as predominant groups (depending on the HP conditions). The application of HTS provided new insights on the diversity and dynamics of the bacterial communities of sous-vide cod, revealing the presence of bacterial genera not previously described in this food, such as Anoxybacillus. The significance of Anoxybacillus as a contaminant of seafoods should be further investigated.
Research – Comparison of Activity of Commercial Protective Cultures and Thermophilic Lactic Acid Bacteria against Listeria monocytogenes: A New Perspective to Improve the Safety of Sardinian PDO Cheeses
Abstract
Listeria monocytogenes contamination that occurs during and post-processing of dairy products is a serious concern for consumers, and bioprotective cultures can be applied to control the growth of the pathogen in sheep milk cheeses. However, to respect specifications provided for protected designation of origin (PDO) cheeses, only autochthonous microorganisms can be used as bioprotective cultures in these products. This in vitro study aimed to evaluate thermophilic lactic acid bacteria (LAB) isolated from sheep milk as bio-preservative agents to control L. monocytogenes growth in PDO cheese. Results were compared with those obtained with a commercial protective culture (cPC) composed of a Lactiplantibacillus plantarum bacteriocin producer designed to inhibit L. monocytogenes growth in cheese. The in vitro antilisterial activities of n.74 autochthonous LAB and a cPC were tested against 51 L. monocytogenes strains using an agar well diffusion assay. In addition, 16S rRNA sequencing of LAB isolates with antilisterial activity was conducted and strains of Lactobacillus helveticus, Lactobacillus delbrueckii subsp. indicus, Lactobacillus delbrueckii subsp. sunkii, Lactobacillus delbrueckii subsp. lactis and Enterococcus faecalis were identified. In this study, 33.6% (74/220) bacterial strains isolated from milk had characteristics compatible with thermophilic LAB, of which 17.6% (13/74) had in vitro antilisterial activity. These results demonstrate that raw sheep milk can be considered an important source of autochthonous thermophilic LAB that can be employed as protective cultures during the manufacturing of Sardinian PDO cheeses to improve their food safety. The use of bioprotective cultures should be seen as an additional procedure useful to improve cheese safety along with the correct application of good hygienic practices during manufacturing and the post-processing stages.
Posted in Decontamination Microbial, Food Micro Blog, Food Microbiology, Food Microbiology Blog, Food Microbiology Research, Food Microbiology Testing, Listeria, Listeria in Cheese, Listeria monocytogenes, microbial contamination, Microbial growth, Microbiological Risk Assessment, Microbiology, Microbiology Investigations, Microbiology Risk, Research
RASFF Alert – Animal Feed – Microbiological Contamination – Hay
Microbiological deviation in hay for animals from Austria in Croatia
RASFF Alert- Animal Feed – Salmonella – Pigs Ear
Salmonella spp. in Ear Pork from Poland in Malta
Research – Surveillance of Vibrio parahaemolyticus pathogens recovered from ready-to-eat foods
This study examined the occurrence of V. parahaemolyticus from ready-to-eat (RTE) food in Delta State, Nigeria. It also characterized antibiotic resistance and virulence gene profile patterns to determine the associated health risk hazard. Food samples total of 380 were collected randomly and assessed for V. parahaemolyticus. V. parahaemolyticus isolates were characterized for their virulence and antibiogram potentials using a phenotypic and polymerase chain reaction (PCR) approach. A total of 42 (11.1%) samples were contaminated with V. parahaemolyticus. In 17/42 (40.5%) of the V. parahaemolyticus-positive samples, the densities were < 10 MPN/g. However, 19/42 (45.2%) and 6/42 (14.3%) of the samples had densities of 10 – 102 and > 102 MPN/g, respectively. A total of 67 V. parahaemolyticus isolates were identified using PCR; 54(80.6%) isolates were multidrug resistant. A total of 22 (32.8%), 39 (58.2%), and 67 (100%) of the V. parahaemolyticus harbored the tdh, trh, and tlh toxin genes, respectively. The T3SS1 gene (vcrD1) was detected in 67 (100%) of the isolates. The T3SS2α genes which were vcrD2, vopB2, and vopT were detected in 21 (31.3%), 11 (16.4%) and 30 (44.8%) of the isolates respectively. Some of the V. parahaemolytics strains harbored the orf8 gene 20 (29.9%), and a combination of orf8 + tdh genes 12 (17.9%), categorized as pandemic strains. The antibiotic resistance genes detected in this study include blaTEM 33 (49.3), tetM 19 (28.4), cmlA 32(47.8) and sul1 14 (20.9). The concentration levels and prevalence of V. parahaemolyticus in RTE foods indicate contamination of ready-to-eat foods, particularly street foods consumed in the Delta State of Nigeria, threatening public health and consumer safety.
Posted in Decontamination Microbial, Food Micro Blog, Food Microbiology, Food Microbiology Blog, Food Microbiology Research, Food Microbiology Testing, microbial contamination, Microbial growth, Microbiological Risk Assessment, Microbiology, Microbiology Investigations, Microbiology Risk, Vibrio, Vibrio parahaemolyticus, Water, water microbiology, Water Safety
Research – High winds can worsen pathogen spread at outdoor chicken farms – Campylobacter
A study of chicken farms in the West found that high winds increased the prevalence of Campylobacter in outdoor flocks, a bacterial pathogen in poultry that is the largest single cause of foodborne illness in the U.S. Researchers found that about 26% of individual chickens had the pathogen at the ‘open environment’ farms in the study, which included organic and free-range chicken farms. High winds the week prior to sampling and the farms’ location in more intensive agricultural settings were linked to a greater prevalence of Campylobacter.
Posted in Campylobacter, campylobacter coli, Campylobacter jejuni, Decontamination Microbial, Food Micro Blog, Food Microbiology, Food Microbiology Blog, Food Microbiology Research, Food Microbiology Testing, microbial contamination, Microbial growth, Microbiological Risk Assessment, Microbiology, Microbiology Investigations, Microbiology Risk


